Project 4173: P. Jałoszyński, X. Luo, R. G. Beutel. 2022. Evolution of cephalic structures in extreme myrmecophiles: a lesson from Clavigeritae (Coleoptera: Staphylinidae: Pselaphinae). Cladistics. 38 (3):335-358.
Abstract
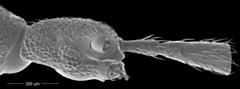
Pselaphinae is a large subfamily, comprising over 10 000 species of the megadiverse Staphylinidae (rove beetles). A remarkable feature of this group is the extreme structural diversity of different body regions, especially the head and its appendages. Within Pselaphinae, Clavigeritae stand out as a clade of highly specialized myrmecophiles. We examined internal and external head structures of the clavigerite species Diartiger kubotai Nomura, using state-of-the-art techniques. The cephalic morphology indicates in a phylogenetic context that the loss of eyes in some Clavigeritae was the latest of major evolutionary changes. We compiled the largest set of morphological data ever scored for the subfamily, comprising 155 characters of the head. Parsimony analyses and Bayesian inference yielded a similar phylogenetic pattern, largely congruent with results published previously. We retrieved Pselaphinae as a clade, and Faronitae as sister to all remaining groups of the subfamily. Faronitae are followed by a “Euplectitae grade” and non-monophyletic Goniaceritae, Batrisitae and Pselaphitae. Clavigeritae are monophyletic, but have evolved within the pselaphite grade. The enigmatic Colilodion Besuchet, recently shifted from Clavigeritae to a paraphyletic Pselaphitae, was placed as sister to extant clavigerites based on an array of cephalic synapomorphies. The current classification of Pselaphinae is unstable and deep changes should be made maintaining only monophyletic units, whereas most of the supertribes are paraphyletic. Characters of the head, with a concentration of mouthparts and sensory structures, and essential parts of the digestive tract and the nervous system, are highly informative phylogenetically. Study of internal structures, presently still at a very preliminary stage, obviously is essential for understanding the evolution of Pselaphinae. Future genetic investigations may reveal mechanisms behind the unique structural megadiversity in this exceptional group of rove beetles.Read the article »
Article DOI: 10.1111/cla.12498
Project DOI: 10.7934/P4173, http://dx.doi.org/10.7934/P4173
| This project contains | Matrices |
|---|---|
Download Project SDD File | Total scored cells: 12523 Total media associated with cells: 0 Total labels associated with cell media: 0 |
| Characters | |
| Total characters: 155 Total characters with associated media: 0 Total characters with media with labels: 0 Total character states: 382 Total character states with associated media: 0 Total character states with media with labels:0 Total unordered/ordered characters:155/0 |
Currently Viewing:
MorphoBank Project 4173
MorphoBank Project 4173
- Creation Date:
21 January 2022 - Publication Date:
11 August 2022 - Project views: 5791

- Matrix downloads: 1

Authors' Institutions ![]()
- Universitaet Jena (FSU Jena)
- University of Wroclaw
Members
| member name | taxa |
specimens |
media | chars | character
| cell scorings (scored, NPA, "-") | cell
| rules | ||||||
| MorphoBank Curator Project Administrator | 82 | 1 | 1 | 155 | 0 | 0 | 12523 (11055, 0, 1468) | 0 | 0 | 0 | ||||
| Pawel Jaloszynski Full membership | 0 | 0 | 0 | 0 | 0 | 0 | 0 (0, 0, 0) | 0 | 0 | 0 | ||||
Taxonomic Overview for Matrix 'M28675' (81 Taxa)
| taxon | unscored cells |
scored cells |
no cell support |
NPA cells |
"-" cells | cell images | labels on cell images |
member access |
| [1] Ocypus olens Last Modified in 01/21/22 | 0 | 130 | 130 | 0 | 25 | 0 | 0 | 2 |
| [2] Dasycerus sulcatus Last Modified in 01/21/22 | 0 | 139 | 139 | 0 | 16 | 0 | 0 | 2 |
| [3] Anthobium atrocephalum Last Modified in 01/21/22 | 0 | 138 | 138 | 0 | 17 | 0 | 0 | 2 |
| [4] Lesteva longoelytrata Last Modified in 01/21/22 | 0 | 141 | 141 | 0 | 14 | 0 | 0 | 2 |
| [5] Omalium rivulare Last Modified in 01/21/22 | 0 | 140 | 140 | 0 | 15 | 0 | 0 | 2 |
| [6] Micropeplus porcatus Last Modified in 01/21/22 | 0 | 137 | 137 | 0 | 18 | 0 | 0 | 2 |
| [7] Megarthrus depressus Last Modified in 01/21/22 | 0 | 136 | 136 | 0 | 19 | 0 | 0 | 2 |
| [8] Metopsia similis Last Modified in 01/21/22 | 0 | 133 | 133 | 0 | 22 | 0 | 0 | 2 |
| [9] Proteinus brachypterus Last Modified in 01/21/22 | 0 | 133 | 133 | 0 | 22 | 0 | 0 | 2 |
| [10] Silphotelus sp. Last Modified in 01/21/22 | 0 | 136 | 136 | 0 | 19 | 0 | 0 | 2 |
| [11] Faronus sp Last Modified in 01/21/22 | 0 | 140 | 140 | 0 | 15 | 0 | 0 | 2 |
| [12] Delenda frivaldszkyi Last Modified in 01/21/22 | 0 | 143 | 143 | 0 | 12 | 0 | 0 | 2 |
| [13] Euplectus karstenii Last Modified in 01/21/22 | 0 | 142 | 142 | 0 | 13 | 0 | 0 | 2 |
| [14] Leptoplectus kijimunaa Last Modified in 01/21/22 | 0 | 143 | 143 | 0 | 12 | 0 | 0 | 2 |
| [15] Plectophloeus nubigena Last Modified in 01/21/22 | 0 | 142 | 142 | 0 | 13 | 0 | 0 | 2 |
| [16] Bibloporus bicolor Last Modified in 01/21/22 | 0 | 141 | 141 | 0 | 14 | 0 | 0 | 2 |
| [17] Bibloplectus ambiguus Last Modified in 01/21/22 | 0 | 144 | 144 | 0 | 11 | 0 | 0 | 2 |
| [18] Philoscotus rostralis Last Modified in 01/21/22 | 0 | 144 | 144 | 0 | 11 | 0 | 0 | 2 |
| [19] Trichonyx sulcicollis Last Modified in 01/21/22 | 0 | 143 | 143 | 0 | 12 | 0 | 0 | 2 |
| [20] Saulcyella schmidtii Last Modified in 01/21/22 | 0 | 144 | 144 | 0 | 11 | 0 | 0 | 2 |
| [21] Trimium brevicorne Last Modified in 01/21/22 | 0 | 144 | 144 | 0 | 11 | 0 | 0 | 2 |
| [22] Batrisodes unisexualis Last Modified in 01/21/22 | 0 | 141 | 141 | 0 | 14 | 0 | 0 | 2 |
| [23] Batrisus formicarius Last Modified in 01/21/22 | 0 | 139 | 139 | 0 | 16 | 0 | 0 | 2 |
| [24] Batrisoplisus torticornis Last Modified in 01/21/22 | 0 | 140 | 140 | 0 | 15 | 0 | 0 | 2 |
| [25] Cratna torticornis Last Modified in 01/21/22 | 0 | 137 | 137 | 0 | 18 | 0 | 0 | 2 |
| [26] Intestinarius quinquesulcatus Last Modified in 01/21/22 | 0 | 142 | 142 | 0 | 13 | 0 | 0 | 2 |
| [27] Mnia elegans Last Modified in 01/21/22 | 0 | 141 | 141 | 0 | 14 | 0 | 0 | 2 |
| [28] Oxarthrius sp. Last Modified in 01/21/22 | 0 | 141 | 141 | 0 | 14 | 0 | 0 | 2 |
| [29] Bergrothia saulcyi Last Modified in 01/21/22 | 0 | 141 | 141 | 0 | 14 | 0 | 0 | 2 |
| [30] Brachygluta fossulata Last Modified in 01/21/22 | 0 | 145 | 145 | 0 | 10 | 0 | 0 | 2 |
| [31] Fagniezia impressa Last Modified in 01/21/22 | 0 | 143 | 143 | 0 | 12 | 0 | 0 | 2 |
| [32] Reichenbachia juncorum Last Modified in 01/21/22 | 0 | 143 | 143 | 0 | 12 | 0 | 0 | 2 |
| [33] Rybaxis longicornis Last Modified in 01/21/22 | 0 | 142 | 142 | 0 | 13 | 0 | 0 | 2 |
| [34] Triomicrus protervus Last Modified in 01/21/22 | 0 | 143 | 143 | 0 | 12 | 0 | 0 | 2 |
| [35] Trissemus antennatus Last Modified in 01/21/22 | 0 | 143 | 143 | 0 | 12 | 0 | 0 | 2 |
| [36] Anasopsis sp. Last Modified in 01/21/22 | 0 | 142 | 142 | 0 | 13 | 0 | 0 | 2 |
| [37] Baraxina sp. Last Modified in 01/21/22 | 0 | 143 | 143 | 0 | 12 | 0 | 0 | 2 |
| [38] Eupines sp. Last Modified in 01/21/22 | 0 | 139 | 139 | 0 | 16 | 0 | 0 | 2 |
| [39] Eupines impar Last Modified in 01/21/22 | 0 | 142 | 142 | 0 | 13 | 0 | 0 | 2 |
| [40] Eupines sphaerica Last Modified in 01/21/22 | 0 | 139 | 139 | 0 | 16 | 0 | 0 | 2 |
| [41] Bryaxis bulbifer Last Modified in 01/21/22 | 0 | 144 | 144 | 0 | 11 | 0 | 0 | 2 |
| [42] Tychobythinus aino Last Modified in 01/21/22 | 0 | 143 | 143 | 0 | 12 | 0 | 0 | 2 |
| [43] Plagiophorus fujiyamai Last Modified in 01/21/22 | 0 | 143 | 143 | 0 | 12 | 0 | 0 | 2 |
| [44] Morana elegans Last Modified in 01/21/22 | 0 | 142 | 142 | 0 | 13 | 0 | 0 | 2 |
| [45] Natypleurus sp. Last Modified in 01/21/22 | 0 | 143 | 143 | 0 | 12 | 0 | 0 | 2 |
| [46] Nedarassus sp. Last Modified in 01/21/22 | 0 | 144 | 144 | 0 | 11 | 0 | 0 | 2 |
| [47] Sunorfa sp. Last Modified in 01/21/22 | 0 | 142 | 142 | 0 | 13 | 0 | 0 | 2 |
| [48] Takaorites torticornis Last Modified in 01/21/22 | 0 | 142 | 142 | 0 | 13 | 0 | 0 | 2 |
| [49] Tenguobythus decoratus Last Modified in 01/21/22 | 0 | 142 | 142 | 0 | 13 | 0 | 0 | 2 |
| [50] Imtempus sp. Last Modified in 01/21/22 | 0 | 144 | 144 | 0 | 11 | 0 | 0 | 2 |
| [51] Pygoxyon sp. Last Modified in 01/21/22 | 0 | 138 | 138 | 0 | 17 | 0 | 0 | 2 |
| [52] Tychus niger Last Modified in 01/21/22 | 0 | 140 | 140 | 0 | 15 | 0 | 0 | 2 |
| [53] Raphitreus speratus Last Modified in 01/21/22 | 0 | 142 | 142 | 0 | 13 | 0 | 0 | 2 |
| [54] Tmesiphoromimus couloni Last Modified in 01/21/22 | 0 | 142 | 142 | 0 | 13 | 0 | 0 | 2 |
| [55] Tmesiphorus crassicornis Last Modified in 01/21/22 | 0 | 140 | 140 | 0 | 15 | 0 | 0 | 2 |
| [56] Ctenistes sp. Last Modified in 01/21/22 | 0 | 136 | 136 | 0 | 19 | 0 | 0 | 2 |
| [57] Pilopius sp. Last Modified in 01/21/22 | 1 | 134 | 134 | 0 | 20 | 0 | 0 | 2 |
| [58] Sognorus breviceps Last Modified in 01/21/22 | 1 | 137 | 137 | 0 | 17 | 0 | 0 | 2 |
| [59] Ancystrocerus sp. Last Modified in 01/21/22 | 0 | 140 | 140 | 0 | 15 | 0 | 0 | 2 |
| [60] Labomimus reitteri Last Modified in 01/21/22 | 0 | 141 | 141 | 0 | 14 | 0 | 0 | 2 |
| [61] Lasinus spinosus Last Modified in 01/21/22 | 0 | 140 | 140 | 0 | 15 | 0 | 0 | 2 |
| [62] Saltisedes kojimai Last Modified in 01/21/22 | 0 | 140 | 140 | 0 | 15 | 0 | 0 | 2 |
| [63] Tyrus mucronatus Last Modified in 01/21/22 | 0 | 141 | 141 | 0 | 14 | 0 | 0 | 2 |
| [64] Pseudophanias sp. Last Modified in 01/21/22 | 0 | 139 | 139 | 0 | 16 | 0 | 0 | 2 |
| [65] Pselaphaulax dresdensis Last Modified in 01/21/22 | 0 | 143 | 143 | 0 | 12 | 0 | 0 | 2 |
| [66] Pselaphogenius treskanus Last Modified in 01/21/22 | 0 | 144 | 144 | 0 | 11 | 0 | 0 | 2 |
| [67] Pselaphus heisei Last Modified in 01/21/22 | 0 | 143 | 143 | 0 | 12 | 0 | 0 | 2 |
| [68] Colilodion Last Modified in 01/21/22 | 1 | 116 | 116 | 0 | 38 | 0 | 0 | 2 |
| [69] Claviger testaceus Last Modified in 01/21/22 | 0 | 117 | 117 | 0 | 38 | 0 | 0 | 2 |
| [70] Claviger longicornis Last Modified in 01/21/22 | 0 | 114 | 114 | 0 | 41 | 0 | 0 | 2 |
| [71] Diartiger kubotai Last Modified in 01/21/22 | 0 | 121 | 121 | 0 | 34 | 0 | 0 | 2 |
| [72] Hadrophorus martinkae Last Modified in 01/21/22 | 3 | 118 | 118 | 0 | 34 | 0 | 0 | 2 |
| [73] Novoclaviger gibbiventris Last Modified in 01/21/22 | 3 | 118 | 118 | 0 | 34 | 0 | 0 | 2 |
| [74] Pararticerus latus Last Modified in 01/21/22 | 3 | 118 | 118 | 0 | 34 | 0 | 0 | 2 |
| [75] Zuluclavodes bryantaylori Last Modified in 01/21/22 | 3 | 118 | 118 | 0 | 34 | 0 | 0 | 2 |
| [76] Disarthricerus integer Last Modified in 01/21/22 | 3 | 113 | 113 | 0 | 39 | 0 | 0 | 2 |
| [77] Tiracaleda minuta Last Modified in 01/21/22 | 3 | 113 | 113 | 0 | 39 | 0 | 0 | 2 |
| [78] Tiracerus sp. Last Modified in 01/21/22 | 3 | 115 | 115 | 0 | 37 | 0 | 0 | 2 |
| [79] Tiraspirus tabulatus Last Modified in 01/21/22 | 2 | 116 | 116 | 0 | 37 | 0 | 0 | 2 |
| [80] Gen.n. neocalexc Last Modified in 01/21/22 | 4 | 113 | 113 | 0 | 38 | 0 | 0 | 2 |
| [81] Cerylambus reticulatus Last Modified in 01/21/22 | 2 | 117 | 117 | 0 | 36 | 0 | 0 | 2 |
Project views 
| type | number of views | Individual items viewed (where applicable) |
| Total project views | 5791 | |
| Project overview | 328 | |
| Views for media list | 326 | |
| Bibliography | 209 | |
| Media views | 990 | Media search (862 views); M838502 (128 views); |
| Documents list | 267 | |
| Taxon list | 2901 | |
| Matrix views | 258 | Matrix landing page (237 views); Data matrix from author (21 views); |
| Specimen list | 512 |
Project downloads 
| type | number of downloads | Individual items downloaded (where applicable) |
| Total downloads from project | 2 | |
| Project downloads | 1 | |
| Matrix downloads | 1 | Data matrix from author (1 download); |