Project 3885: L. L. Voyta, A. V. Abramov, L. A. Lavrenchenko, V. Nicolas, E. A. Petrova, L. Y. Kryuchkova. 2021. Dental polymorphisms in Crocidura (Soricomorpha: Soricidae) and evolutionary diversification of crocidurine shrew dentition. Zoological Journal of the Linnean Society. Early View:null.
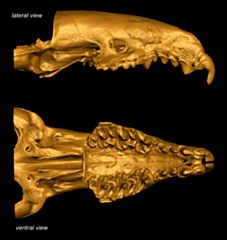
Specimen: Crocidura yaldeni Lavrenchenko, Voyta, Hutterer, 2016 (ZMMU/Theriology:S-165343)
View: 2D surface of upper teeth
View: 2D surface of upper teeth
Abstract
The upper dentition of Crocidura exhibits polymorphic characters that were revealed for the first time in this study via high-resolution X-ray computed microtomography. Our analyses of 11 Crocidura species and selected Diplomesodon, Suncus and Sylvisorex species from different geographical regions and size groups revealed the most complex character states of upper dentition in the Ethiopian endemic species Crocidura yaldeni. A three-dimensionally based geometric morphometric analysis revealed the dependence of variation in skull muzzle shape on alterations in general upper dentition, such as a reduction in the number of antemolars. Principal components analysis revealed highly significant shape alterations and morphological trajectories in C. yaldeni (and more moderate ones in Suncus murinus) toward the Sorex-like morphotype in the outgroup, and less significant shape alterations in Crocidura obscurior, Crocidura phanluongi and Crocidura sapaensis with double-rooted third antemolar. Cladistic analysis based on a new data matrix for 20 species and 46 characters allowed us to determine the directions of the morphological trajectories: the apomorphic state of the most complex antemolars of C. yaldeni is associated with deviating skull muzzle shape changes, which we determined to be attributable to neomorphosis, and the less significant alterations in the shape of other Crocidura with complex antemolars are attributable to regional adaptation.Read the article »
Article DOI: 10.1093/zoolinnean/zlab103
Project DOI: 10.7934/P3885, http://dx.doi.org/10.7934/P3885
| This project contains | Matrices |
|---|---|
Download Project SDD File | Total scored cells: 917 Total media associated with cells: 1 Total labels associated with cell media: 0 |
| Characters | |
| Total characters: 46 Total characters with associated media: 46 Total characters with media with labels: 0 Total character states: 108 Total character states with associated media: 0 Total character states with media with labels:0 Total unordered/ordered characters:45/1 |
Currently Viewing:
MorphoBank Project 3885
MorphoBank Project 3885
- Creation Date:
24 November 2020 - Publication Date:
21 December 2021 - Media downloads: 8

- Matrix downloads: 9

Authors' Institutions ![]()
- Muséum national d'Histoire naturelle, Paris (Natural History Museum, Paris)
- Centre National de la Recherche Scientifique (CNRS)
- Saint Petersburg State University
- Sorbonne Université
- Russian Academy of Sciences
- Severtsov Institute of Ecology and Evolution
- Université des Antilles
- Zoological Institute of the Russian Academy of Sciences
Members
| member name | taxa |
specimens |
media | media notes | chars | character
| cell scorings (scored, NPA, "-") | cell
| rules | ||||||||||||||
| Leonid Voyta Project Administrator | 21 | 36 | 186 | 186 | 46 | 0 | 46 | 46 | 0 | 917 (860, 0, 57) | 0 | 0 | 1 | 0 | 0 | ||||||||
Taxonomic Overview for Matrix 'M27141' (20 Taxa)
| taxon | unscored cells |
scored cells |
no cell support |
NPA cells |
"-" cells | cell images | labels on cell images |
member access |
| [1] Sorex minutissimus Taxon name last Modified on 12/20/20 | 0 | 44 | 44 | 0 | 2 | 0 | 0 | 1 |
| [2] Sorex mirabilis Taxon name last Modified on 12/20/20 | 0 | 46 | 46 | 0 | 0 | 0 | 0 | 1 |
| [3] Neomys fodiens Taxon name last Modified on 12/20/20 | 0 | 41 | 41 | 0 | 5 | 0 | 0 | 1 |
| [4] Myosorex okuensis Taxon name last Modified on 12/20/20 | 2 | 41 | 41 | 0 | 3 | 0 | 0 | 1 |
| [5] Suncus etruscus Taxon name last Modified on 12/20/20 | 0 | 44 | 43 | 0 | 2 | 1 | 0 | 1 |
| [6] Suncus megalura Taxon name last Modified on 12/20/20 | 0 | 45 | 45 | 0 | 1 | 0 | 0 | 1 |
| [7] Suncus murinus Taxon name last Modified on 12/20/20 | 1 | 41 | 41 | 0 | 4 | 0 | 0 | 1 |
| [8] Sylvisorex johnstoni Taxon name last Modified on 12/20/20 | 0 | 42 | 42 | 0 | 4 | 0 | 0 | 1 |
| [9] Crocidura jouvenetae Taxon name last Modified on 12/20/20 | 0 | 45 | 45 | 0 | 1 | 0 | 0 | 1 |
| [10] Crocidura olivieri Taxon name last Modified on 12/20/20 | 0 | 44 | 44 | 0 | 2 | 0 | 0 | 1 |
| [11] Crocidura obscurior Taxon name last Modified on 12/20/20 | 0 | 42 | 42 | 0 | 4 | 0 | 0 | 1 |
| [12] Crocidura yaldeni Taxon name last Modified on 12/20/20 | 0 | 44 | 44 | 0 | 2 | 0 | 0 | 1 |
| [13] Crocidura lasiura Taxon name last Modified on 12/20/20 | 0 | 43 | 43 | 0 | 3 | 0 | 0 | 1 |
| [14] Crocidura leucodon Taxon name last Modified on 12/20/20 | 0 | 43 | 43 | 0 | 3 | 0 | 0 | 1 |
| [15] Crocidura serezkyensis Taxon name last Modified on 12/20/20 | 0 | 44 | 44 | 0 | 2 | 0 | 0 | 1 |
| [16] Crocidura zarudnyi Taxon name last Modified on 12/20/20 | 0 | 45 | 45 | 0 | 1 | 0 | 0 | 1 |
| [17] Crocidura phanluongi Taxon name last Modified on 12/20/20 | 0 | 42 | 42 | 0 | 4 | 0 | 0 | 1 |
| [18] Crocidura sapaensis Taxon name last Modified on 12/20/20 | 0 | 43 | 43 | 0 | 3 | 0 | 0 | 1 |
| [19] Crocidura zaitsevi Taxon name last Modified on 12/20/20 | 0 | 45 | 45 | 0 | 1 | 0 | 0 | 1 |
| [20] Diplomesodon pulchellum Taxon name last Modified on 12/20/20 | 0 | 36 | 36 | 0 | 10 | 0 | 0 | 1 |
Project downloads 
| type | number of downloads | Individual items downloaded (where applicable) |
| Total downloads from project | 51 | |
| Media downloads | 8 | M737978 (1 download); M738402 (1 download); M738415 (1 download); M738416 (1 download); M738417 (1 download); M738418 (1 download); M737980 (1 download); M738436 (1 download); |
| Project downloads | 34 | |
| Matrix downloads | 9 | Crocidurinae matrix (v. 03) (9 downloads); |
